If checked (unchecked is the default), distances in the file to be converted are assumed to be in Bohr rather than angstroms ( -ubohr). The radio buttons on this window specify the format of the file to be converted: Radio ButtonĮxtract a Z-matrix from a Gaussian 16 checkpoint file, from the optimization step indicated in the Checkpt Geom Opt. This option is not implemented in the current version of Gaussian 16W. If checked, edit the new input file after conversion using the designated external editor (default is unchecked). If checked, edit the new input file after converting using the Job Edit window (default is unchecked). The default is checked if the selected file is not a. If checked, load the converted job into memory. The Load button (appears as a file folder icon) can be used to load a different input file to be converted. The Cancel button returns you to the Job Processing window without converting any file. The Convert button begins the conversion of the designated file, using currently set options. The default is the filename of the file to be converted with the extension GJF. The Generated Filename field is used to specify the name of the converted file. It is an interface to the standard NewZMat utility which is included with Gaussian 16. By default, it converts files to Gaussian 16W input files (.GJF files). It is reached via the NewZMat item on the Utilities menu. The NewZMat File Conversionwindow controls file conversions. We will consider them here in the same order as they appear in that menu, beginning with the NewZMat facility. They are accessible via the various items on the Utilities menu in the program’s main window. The bond vector definition is here: def get_bond_vector(r, a, t):Īgain, the only part I don't understand is # get local axis system from 3 coordinates.A number of utility programs are included with Gaussian 16W. 
Here is some context of how this function is used: bond_vector = get_bond_vector(atom.rval, atom.aval, al)ĭisp_vector = np.array(np.dot(bond_vector, _axes))Ītom.coords = + disp_vector 
What is "getting local axis system from 3 coordinates"? U23c21 = get_ucp(u23, u21) # unit cross product Print('\nError: Co-linear atoms in an internal coordinate definition') U23 = get_u12(coords2, coords3) #calculating vector between that points 2-3 U21 = get_u12(coords1, coords2) #calculating vector between that points 1-2 However, in his work, there is a mathematical part that I don't understand: # get local axis system from 3 coordinatesĭef get_local_axes(coords1, coords2, coords3): While searching I found TMPChem's work on GitHub and it does exactly what I want. If there aren't any mistakes up to this point, how I will calculate Cartesian coordinates of the 4th atom using these $x,y,z$ values? To calculate $x,y,z$ values from the formulas x = r * sin(theta) * cos(phi) I am not sure if I should calculate this as z2 + s(angle) or z2 - s(angle) and what it depends on, if both are possible.įor the 4th atom, I use spherical coordinates r, theta, phi = (0.976, 96.572, -179.995) 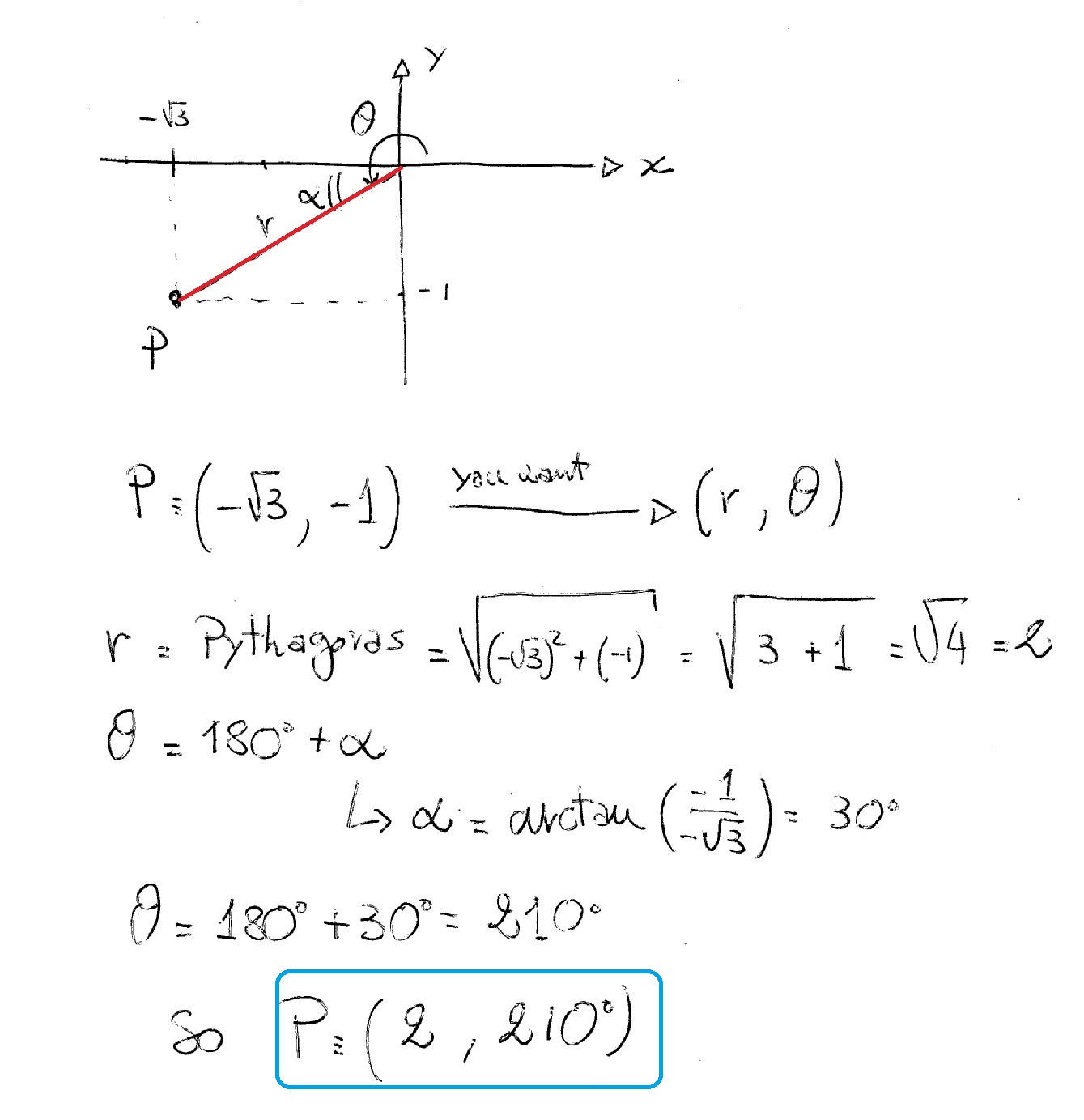
The 3rd atom must have coordinates that are something like this if read correctly: 3 H 0 distance*sin(angle) z2+distance*cos(angle) How should I treat the 3rd and 4th atoms? My question is after setting first atom as 0,0,0 1 O 0 0 0Īnd the second one as 0,0,(distance from first) to put it on the z-axis 2 O 0 0 1.45335189476 This is converting a Z-matrix to Cartesian coordinates. Now I need to perform the reverse operation and use this Z-matrix as input and define $x,y,z$ coordinates for each atom. Then, the script constructs a Z-matrix with them, like this: Z-mat : These are $x,y,z$ coordinates of H2O2 molecule. To begin with, I wrote a script that gets Cartesian coordinates of molecule as input in the below. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
